In 2002, none of the guests at a hotel realized that they had all become hosts or carriers of an unfamiliar virus. In just six months, the deadly grip of that virus claimed 774 lives. The economy was devastated. According to the Asian Development Bank, the loss amounted to 59 billion US dollars. The virus was named “Severe Acute Respiratory Syndrome Coronavirus” (SARS-CoV) because the disease was identified as “Severe Acute Respiratory Syndrome”. But it didn’t end there. A few years later, Saudi Arabia faced the outbreak of “Middle East Respiratory Syndrome Coronavirus” (MERS-CoV). It was found that the MERS-CoV virus was related to SARS-CoV, and panic quickly spread because the world had already witnessed the horrors of SARS-CoV. Although later “epidemiological reports” confirmed that the infection rate of respiratory disease caused by MERS-CoV was lower than that of SARS-CoV, MERS-CoV still had a considerable negative impact worldwide. Many restrictions were imposed on travel between affected and unaffected regions of the world.
Late December, 2019, Wuhan City, China:
People began knocking on doctors’ doors with health complaints—symptoms like fever, cough, shortness of breath, and muscle pain. Doctors reported, “Viral bilateral pneumonia” (that is, both lungs affected by a virus-induced pneumonia). However, alarm soon set in among the doctors as the number of sick people soared. Delving deep into the root cause of the illness, doctors discovered that, masked as pneumonia, a new disease was making its way from the patients’ lungs to their gastrointestinal organs. Nearly seventeen years after humanity suffered the deadly attack of the SARS-CoV virus in 2002, another dangerous virus—previously unknown to humans—was found in the respiratory tracts of people once again.
It was revealed that, taxonomically, the virus belonged to the order “Nidovirales”. Although its genetic material is “RNA” (Ribonucleic Acid), unlike many well-known RNA viruses (such as HIV or Human Immunodeficiency Virus), it does not convert its RNA into “DNA” (Deoxyribonucleic Acid) and integrate into human cellular DNA. Instead, after entering the human cell, this virus disables the cell’s own “messenger RNA” (mRNA) and installs its own RNA in the role of messenger RNA. The cell’s ribosomes, unaware, follow the nucleotide sequence of this viral “messenger RNA” and produce two types of polyproteins (polyproteins are long molecular chains formed by connecting several smaller protein molecules with covalent bonds). Among these, one type of polyprotein breaks down to form sixteen types of “nonstructural proteins” (proteins that assist in the functional, not structural, aspects of the cell), some of which copy the viral RNA via “transcription” (the process of copying the sequence of nucleotides in viral RNA) and “replication” (the process of making new RNA based on the copied sequence), thereby producing many exact copies of RNA necessary for the birth of new viruses—in other words, the virus easily paves the way for its own proliferation inside human cells.
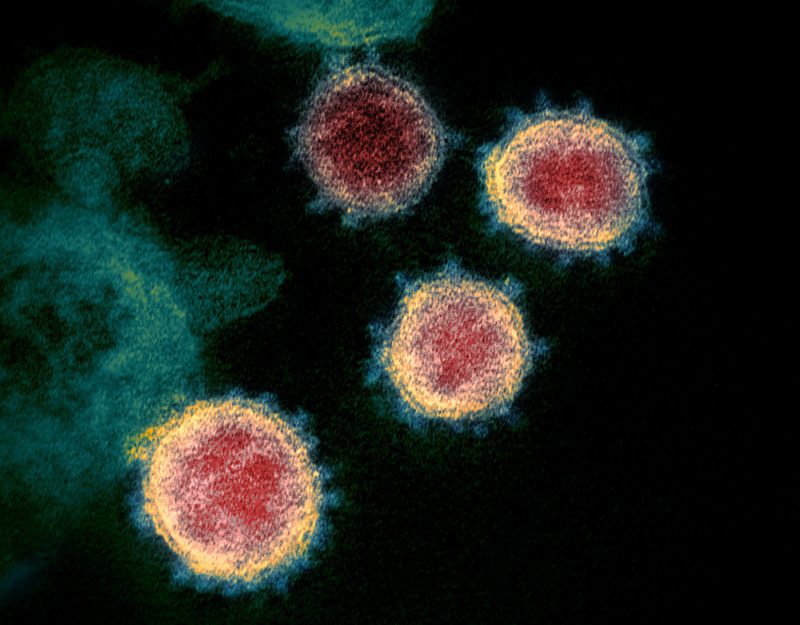
The new contagious disease caused by this virus was named “Coronavirus Disease – 2019” or “COVID-19,” which has unleashed a wave of deaths worldwide and driven the global economy to the brink. Of the four major proteins that make up the virus particle, the most important is the “spike protein.” It’s called the “spike protein” because it forms numerous spike-like projections all over the virus, appearing under a microscope like the “corona” or aura around a bright sun, hence the name “corona.”
This appearance confirmed to scientists that the virus belonged to the “Coronaviridae” family. As this was its first known entry into human civilization, it was initially named the “novel coronavirus”. However, because its RNA nucleotide sequence was about seventy percent similar to that of the SARS-CoV virus, and because it infected the same hosts (such as humans) in similar ways as SARS-CoV, it was later officially named “SARS-CoV-2” or “Severe Acute Respiratory Syndrome Coronavirus-2”. According to viral taxonomy, this virus belongs to the Coronaviridae family, which comprises four genera: “Alphacoronavirus”, “Betacoronavirus”, “Gammacoronavirus”, and “Deltacoronavirus”. Its “identity card” genus is Betacoronavirus. The subgenus is “Sarbecovirus” and the species is “Severe Acute Respiratory Syndrome-related Coronavirus”.
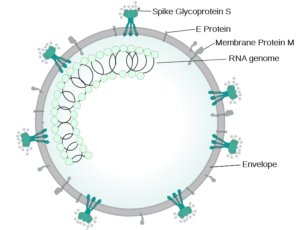
Structure of SARS-CoV-2 Virion Particle and Its Entry into Human Cells:
The diameter of a SARS-CoV-2 virion particle (a virus particle is called a “virion” when it is outside a host body or between two host cells. Once inside the host cell, it’s called a “virus”) ranges from 50 to 200 nanometers. Like other betacoronaviruses, the SARS-CoV-2 particle is constructed from four structural proteins: “spike protein,” “envelope protein,” “membrane protein,” and “nucleocapsid protein.” The “nucleocapsid protein” holds the viral RNA, while the other three proteins form the outer envelope of the virion.
SARS-CoV-2 is particularly inclined to bind to the angiotensin converting enzyme–2 (ACE2) on human cells. This enzyme is abundant in type-2 alveolar cells of the human lungs. It is also plentiful in the glandular cells of the gastric, duodenal, and rectal epithelium, as well as in the endothelial cells and enterocytes of the small intestine. The SARS-CoV-2 virion uses its envelope’s spike protein to attach to the ACE2 enzyme on the human cell and then mainly enters the cell in two ways—
a) The virion fuses its envelope with the “endosome” membrane inside the human cell and enters the “endosome” (an endosome is a membrane-bound compartment inside a eukaryotic cell).
Or
b) The virion fuses its envelope directly with the human cell membrane and enters the cell directly.
Though this is an RNA virus, unlike many familiar RNA viruses (such as HIV), it does not convert its RNA into DNA and integrate into human genome. Instead, after entering the human cell, the virus disables the cell’s own messenger RNA and substitutes its own RNA in its place for the cellular “messenger RNA” function. The human cell’s ribosome, unsuspecting, follows the nucleotide sequence of this viral “messenger RNA” and produces two types of polyprotein. One of these polyproteins breaks down to form sixteen kinds of “nonstructural proteins,” some of which copy the viral RNA via “transcription” and “replication”, generating numerous identical viral RNAs to create the next generation of viruses.
Symptoms of COVID-19:
So far, reported COVID-19 cases show the full spectrum—from minor symptoms and severe illness to death. We know that the time between exposure to a harmful germ, chemical, or radiation and appearance of symptoms is called the “incubation period.” For SARS-CoV-2, the incubation period is about the same as for MERS-CoV, which is two to 14 days. Notable symptoms include—
- Fever
- Cough
- Shortness of breath
- Muscle pain
Prodromal Signs of COVID-19:
Some prodromal signs may appear even before the incubation period begins after SARS-CoV-2 infection, such as—
- Persistent chest pain
- Bluish tint on face or lips
Who is at Higher Risk of Severe COVID-19 Illness:
COVID-19 is a new disease. We still do not have complete information about the risk factors besides possible symptoms and prodromal signs. Those at greater risk of severe illness include—
- People aged 65 or older
- Individuals with lung diseases or problems such as asthma
- People suffering from heart conditions
- Those whose immunity is weakened due to cancer treatment, bone marrow (bone marrow) transplantation, uncontrolled AIDS, or long-term corticosteroid use
- While it’s not yet clear whether pregnant women are at extra risk for COVID-19 complications, in general, they tend to get sicker with any viral infection and thus require close observation
How does COVID-19 spread?
There is ongoing debate as to whether the SARS-CoV-2 virus arose naturally—through genetic mutation of a previously existing virus’s RNA nucleotide sequence—or whether it is artificial, created through human “gene editing” of viral RNA (in other words, if it is a “genetically engineered” virus).
When members of a specific species or subspecies serve as a reservoir for a particular pathogen (infectious virus, bacteria, or other germ) in large numbers, that population is called the pathogen’s “reservoir population”. If that “reservoir population” comes into contact with a “novel host population” (another species or subspecies not previously exposed), the pathogen can cross over. The process by which a pathogen is transmitted from one species or subspecies to another is called “spillover infection”. After entering a “novel host population”, the pathogen may or may not spread extensively.
The “reservoir population” of this virus has yet to be definitively identified, but it is thought that animals became “novel hosts” of the virus through spillover infection, and then human infection began via animal-to-human transmission.
Human-to-human transmission
If you are within six feet of an infected person, droplets expelled from their nose or mouth during coughing or sneezing carry the virion and can land on a healthy person’s nose, mouth, or eyes, facilitating person-to-person transmission.
Transmission from objects or surfaces contaminated with SARS-CoV-2 virions
The properties of the surrounding environment—such as pH (which measures how acidic or alkaline a solution is), voltage, exposure to chromophores (molecular groups responsible for giving color), light, ion (charged atomic or molecular species) concentration, phosphorylation (addition of a phosphate group during a chemical reaction), or the binding of a ligand to form a large molecular-weight complex—can change the shape of large molecules like proteins, sometimes leading to deactivation.
As already mentioned, the envelope of this virion is made up of three proteins: spike, envelope, and membrane protein. When SARS-CoV-2 virions remain outside a host for a long period, envelope proteins may be deactivated by environmental factors, thus reducing the virion’s ability to enter host cells. Experimental observations show that these virions remain active for 18 hours on copper, 55 hours on cardboard, 90 hours on stainless steel, and over 100 hours on plastic. SARS-CoV-2 virions remain active for three hours as aerosols. The virus has also been detected in human and animal feces.
If the envelope of SARS-CoV-2 virions on a surface remains active, touching these objects or surfaces and then touching one’s nose, mouth, or eyes can lead to infection.
Possible Approaches for Treating COVID-19:
We have already received various guidelines for avoiding SARS-CoV-2 virions. But what about paths for counteracting the virus’s harmful effects after it enters the body—that is, a possible way to cure COVID-19? We know that SARS-CoV-2 belongs to the species “Severe Acute Respiratory Syndrome-related Coronavirus”. I propose a possible approach that, if successful, could potentially inhibit the proliferation of not only SARS-CoV-2 but all viruses of this species within human cells, which could enable the cure of COVID-19 and similar diseases. First, let’s briefly review how these viruses multiply in humans—
Replication Method of Severe Acute Respiratory Syndrome-related Coronavirus in Humans
After a virion enters a human cell, the viral nucleocapsid is released into the cytoplasm, and its RNA is freed from the nucleocapsid. The virus disables the cell’s own messenger RNA. The cell’s ribosomes, recognizing only the viral RNA as active, translate, through two-thirds of its full length, two large polyprotein molecules—PP1A and PP1AB.
Each polyprotein chain contains three enzyme molecules: “protease”, “PLpro” (papain-like proteinase), and “3CLpro”, all of which assist in breaking down the polyproteins. The PP1AB polyprotein, when broken down, produces sixteen nonstructural proteins—NSP1, NSP2, …, NSP16. Among the nonstructural proteins, some act as “replication proteins” and are called “nonstructural replication proteins”. These, in combination, form a multi-protein “replicase-transcriptase complex”. The primary “replicase-transcriptase” protein in this complex is the “RNA-dependent RNA polymerase” (NSP12), which plays a key role in viral transcription and replication. The rest of the nonstructural proteins in the complex also assist with transcription and replication. It is through these processes that the viral RNA is faithfully copied and new viral RNA is assembled for the next generation.
It should be noted that the “NSP1” protein and “papain-like proteinase” enzyme play important roles in weakening the human immune system. Additionally, NSP1 is a key destroyer of the cell’s native messenger RNA, thereby hindering the cell’s normal translation process.
Inhibiting Papain-like Proteinase Produced by Severe Acute Respiratory Syndrome-related Coronavirus in Human Cells
Papain enzyme is mainly derived from raw papaya and is used in various food and pharmaceutical industries—for example, clarifying cloudy lenses, tenderizing meat, lysing wound adhesions, and treating stings from hymenoptera and jellyfish. It is also used as an additive in laxatives, tooth powders, digestive aids, and skin lotions. Many people employed in these industries come in contact with papain enzyme and must endure symptoms such as breathing difficulty, chest pain, and cough—symptoms that overlap with those of COVID-19. Therefore, the presence of viral “papain-like proteinase” enzyme in the infected cells of COVID-19 patients may contribute to breathing problems, chest pain, and coughing. If we can use an anti-enzyme (an agent—especially an antibody or enzyme that inhibits the action of another enzyme) against papain-like proteinase in the bodies of COVID-19 patients, it may inhibit the effect of the papain-like enzyme, thereby preventing the breakdown of PP1AB polyprotein and the subsequent formation of nonstructural proteins, which could halt the proliferation of the virus. Employees in papain-dependent industries may have developed such anti-enzymes in their bodies from prolonged exposure, but this needs to be tested and investigated.










good
Thank you.
ভাইয়া, আপনার এই লিখাটি পড়ে করোনা ভাইরাস এর অনেক কথা জানলাম
সবমিলিয়ে ভালই মনে হয়েছে। পোস্টটির জন্য আপনাকে অসংখ্য ধন্যবাদ।
ধন্যবাদ
এই রকম মহামারী নতুন নয়। যুগে যুগে অনেক মহামারী এসেছে আবারও চলে গেছে
ঠিক